|
Biomarkers such as Marker Of Proliferation Ki-67 ( Ki-67), p 16IN4a, a tumor suppressor protein in humans encoded by CDKN2A gene and BDProExC, a recently developed immunocytochemical assay that targets the expression of topoisomerase II-alpha and minichromosome maintenance protein-2 7 have been suggested as biomarkers for improving the clinical performance of cervical cancer screening 3. Utilization of biomarkers in cervical histology and cytological examination has been shown to overcome false positive and false negative issues. So, earmarking the overlapping genes that express differentially at late stages of neoplasia and cancer may be a better approach. Early stages of neoplasia have minimal cytological and histological changes and mostly revert back to normal state on their own. Most of the techniques utilized for detection of cervical cancer are visual in nature with cervicography being fairly common 5, 6. However, the Pap test is entirely dependent on manual cytological screening and visualization of de-shaped, transformed and altered cervical cells, resulting in high false negative and false positive rates 4. Papanicolaou test, also known as Pap smear test is mostly employed for the screening and diagnosing of cervical neoplasia cells 3. Cervical cancer is preceded by a long phase of morphological alteration in cervical cells known as cervical intra-epithelial neoplasia (CIN), which is further characterized as mild (CIN1), moderate (CIN2) and severe (CIN3) cervical dysplasia and finally leading to cervical cancer. Most cases of cervical cancer are caused due to infection with human papillomavirus (HPV) 2. Capitalizing upon and targeting Minichromosome maintenance protein complex - specifically the MCM2, MCM4 and MCM6 subunits, ZWINT and CDC7 for experimental validation, may provide valuable insights in understanding and detection of progressing cervical neoplasia to cervical cancer at an early stage.Ĭervical cancer has been reported to be the second deadliest cancer in women worldwide 1. This overlapping relationship provides a list of common disease related genes among pre-cancerous and cancer stages which could help in targeting the proliferating cancerous cells during onset. Of these, ZWINT, CDC7, MCM4, MCM2 and MCM6 were further identified to be part of most significant motifs. DEGs namely NDC80, ZWINT, CDC7, MCM4, MCM2 and MCM6 were found to be common in both the significant modules as well as the hubs. 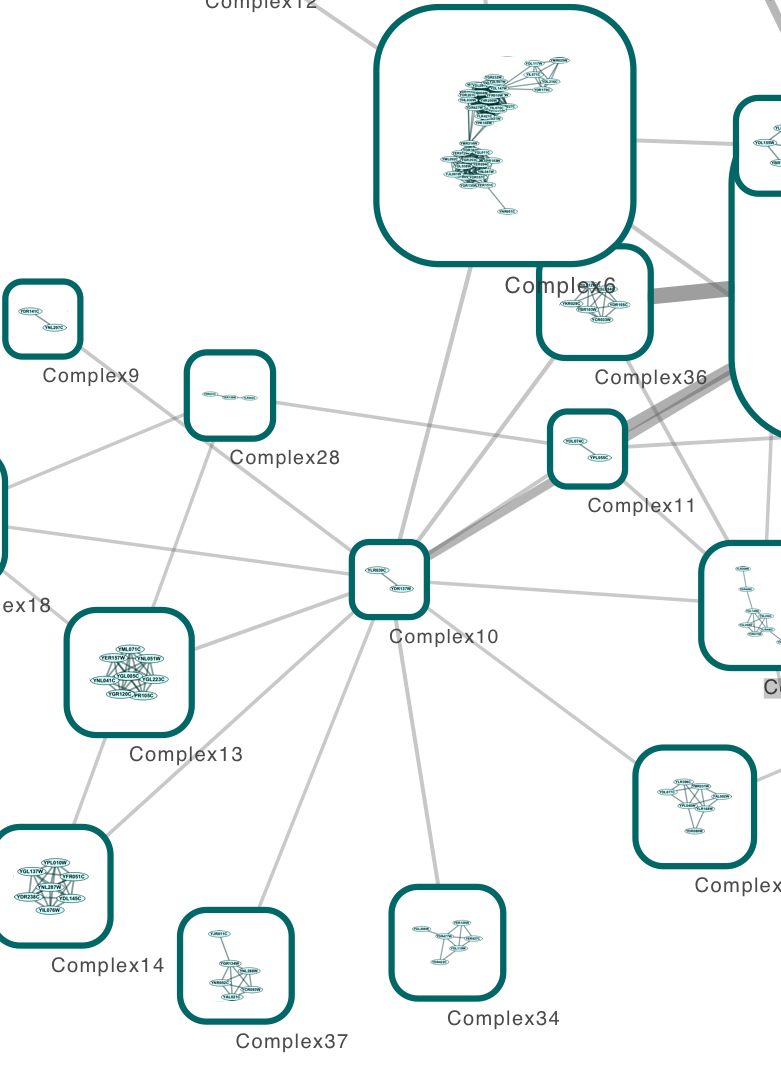
Two significant modules and 10 hubs of the common gene interaction network, with 125 nodes and 201 edges were found. Gene interaction network was constructed using the 98 common DEGs among the dysplastic and cancer cells and analysed for the identification of common modules, hubs and significant motifs. Differentially expressed genes (DEGs) of dysplastic (CIN2 and CIN3) and cancer cells were identified by microarray data analysis. Overlapping genes across high-grade squamous intraepithelial lesions (CIN2 and 3) and cancer may serve as potential biomarkers for this progressive disease.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
